TAGMe DNA Methylation Detection Kits(qPCR) for Cervical Cancer
PRODUCT FEATURES
Precision

Validated over 36000 clinical samples in double-blind multi-center studies, the product has a specificity of 94.3% and a sensitivity of 96.0%.
Convenient

The original Me-qPCR methylation detection technology can be completed in one step within 3 hours without bisulfite transformation.
Early

Cervical cancer screening can be advanced to the stage of high-level lesions (precancerous lesions) stage.
Automation

Accompanied by customized result analysis software, the interpretation of the results is automated and directly readable.
INTENDED USE
This kit is used for in vitro qualitative detection of hypermethylation of the gene PCDHGB7 in cervical specimens. A positive result indicates an increased risk of grade 2 or higher-grade/more advanced cervical intraepithelial neoplasia (CIN2+, including CIN2, CIN3, adenocarcinoma in situ, and cervical cancer), which requires further colposcopy and/or histopathological examination. On the contrary, negative test results indicate that the risk of CIN2+ is low, but the risk cannot be completely excluded. Final diagnosis should be based on colposcopy and/or histopathological results. PCDHGB7 is a member of protocadherin family γ gene cluster. Protocadherin has been found to regulate biological processes such as cell proliferation, cell cycle, apoptosis, invasion, migration and autophagy of tumor cells through various signaling pathways, and its gene silencing caused by hypermethylation of the promoter region is closely related to the occurrence and development of many cancers. It has been reported that hypermethylation of PCDHGB7 is associated with a variety of tumors, such as non-Hodgkin lymphoma, breast cancer, cervical cancer, endometrial cancer and bladder cancer.
DETECTION PRINCIPLE
This kit contains nucleic acid extraction reagent and PCR detection reagent. Nucleic acid is extracted by magnetic-bead-based method. This kit is based on the principle of fluorescence quantitative PCR method, using methylation-specific real-time PCR reaction to analyze template DNA, and simultaneously detect the CpG sites of PCDHGB7 gene and the quality control marker internal reference gene fragments G1 and G2. The methylation level of PCDHGB7 in the sample, or the Me value, is calculated according to the PCDHGB7 gene methylated DNA amplification Ct value and the Ct value of the reference. The PCDHGB7 gene hypermethylation positive or negative status is determined according to the Me value.

Application scenarios
Early screening
Healthy people
Cancer Risk Assessment
High-risk population (positive for high-risk human papillomavirus (hrHPV) or positive for cervical exfoliation cytology)
Recurrence Monitoring
Postoperative population (with a history of high-grade cervical lesions or cervical cancer)
Clinical significance
Early screening for healthy population: Cervical cancer and precancerous lesions can be accurately screened out
Risk assessment in high-risk population: Risk classification can be performed in HPV-positive populations to guide subsequent triage detection
Recurrence monitoring for postoperative population: Postoperative population recurrence monitoring can be performed to prevent delays in treatment caused by recurrence
Sample collection
Sampling method: Place the disposable cervical sampler at the cervical os, gently rub the cervical brush and rotate 4-5times clockwise, slowly remove the cervical brush, put it into cell preservation solution, and label it for the following examination.
Preservation of samples: Samples can be stored at room temperature for up to 14 days, at 2-8 ℃ for up to 2 months, and at -20±5℃ for up to 24 months.
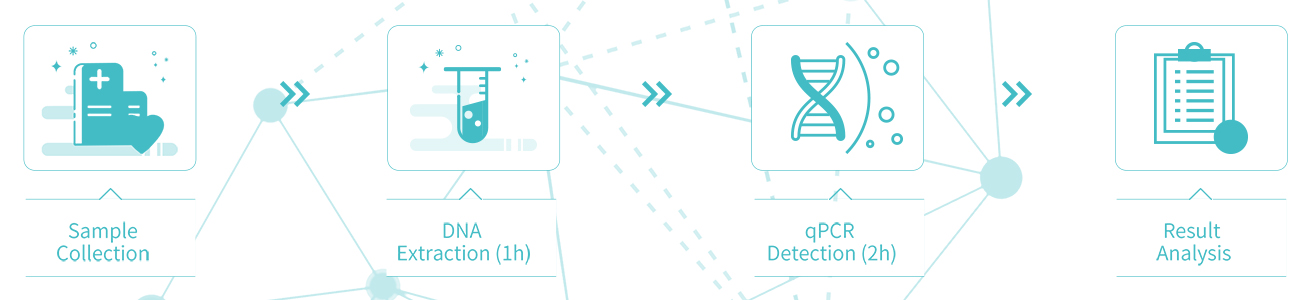
Detection process:3 Hours (Without manual process)
DNA Methylation Detection Kits (qPCR) for Urothelial Cancer

|
Clinical application |
Clinical auxiliary diagnosis of cervical cancer |
|
Detection gene |
PCDHGB7 |
|
Sample type |
Female cervical specimens |
|
Test method |
Fluorescence quantitative PCR technology |
|
Applicable model |
ABI7500 |
|
Packing specification |
48 tests/kit |
|
Storage Conditions |
Kit A should be stored at 2-30℃
Kit B should be stored at -20±5℃ Valid for up to 12 months |